If you've ever folded a paper airplane, chances are you folded The Dart. It's fast and simple, and kids all around the world have launched it through the air. Soon, however, they will move on to other designs – airplanes that glide, make loops, or twirl in flight. There are thousands of paper airplane designs, each with its own quirks and structure.

In the artificial life world, we would call paper airplane design a quality-diversity algorithm. A paper airplane should be fly fast, but we don’t only care about an optimal design – we’re also interested in airplanes with unique bodies or wing types. Rather than a single design, quality-diversity algorithms aim to develop a population of diverse solutions to a problem.
In this post, let’s explore how quality-diversity algorithms can give us a peek into a world of swimming, soft creatures.
Oceanworld
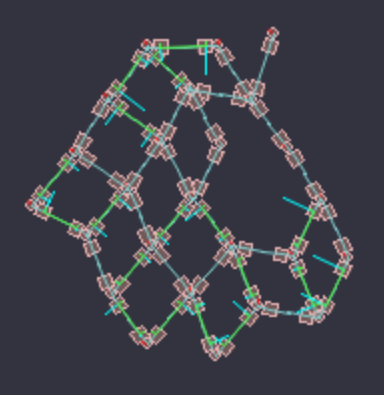
A creature moves by a periodic heartbeat. Whenever a heartbeat begins, any muscle bonds will extend their length, deforming the shape of the creature. When the heartbeat ends, the muscle bonds will return to original length. In addition, a bond can also be rigid. Rigid bonds are affected by fluid resistance – when they move, a force will pushback in the opposite direction.
The great part about evolutionary methods is that this is all a black box -- we don't actually need to understand how the physics works, we just need to run the algorithm.
Cool, we have some creatures. But how much of the total possible space do these creatures actually represent?
One way we can gain intuition about the possible space is through a technique called novelty search. Novelty search tries to find a population of creatures that are diverse from each other according to some metric. In our case, let’s use the number of nodes, and the ratio of bonds to nodes.
Novelty Search works by a simple loop:
- Maintain a population of 1000 genomes.
- Mutate each genome and add it to the population.
- Sum the distance each genome has to its 10 closest neighbors, in terms of some novelty metric.
- Sort the populations by increasing distance, and keep only the top 1000.
Working correctly, the novelty search algorithm should optimize for a group of genomes that are spread out in terms of number of nodes and bonds-node ratio.
I always like to sanity check by looking in, so let’s take a peek at some sample points and see how the creatures look.
There’s a key thing missing in our Oceanworld creatures – they don’t actually do anything. The “Quality” in Quality-Diversity has to measure something. So let’s choose something: a creature that swims faster is better.
First, let’s try out a greedy evolutionary algorithm. We’ll start with a random creature, and measure how far it travels. Then, we’ll randomly mutate its genome and try again. If the new creature performs better, it replaces the old one.
Like a pair of deformed jellyfish, our creatures have evolved to move forwards by pulsating their muscle bonds. I doubt I would have been able to hand-design any of these creatures – the physics relationships are just too complex. Thankfully, evolution takes care of it for us.
But are these really the best creatures? The key observation here is that different starting genomes converge to different solutions. This tells us that our possibility space has local minima – genomes where any small change will hurt performance, even though a larger change could be beneficial.
It turns out that local minima are part of a classic paradox: “the path to success does not look like success”. When Carl Benz proposed the motor engine in a world of horse carriages, it was slow and heavy. But he kept at it, and motor cars emerged the victor.
In some form, Benz was using a quality-diversity algorithm: motor engines were different, so they deserved a chance. Quality-diversity algorithms help us escape local minima, because they force us to consider a group of genomes rather than a single leader. With more diversity, there is more chance to discover a path forwards.
MAP-Elites
MAP-Elites is a quality-diveristy algorithm that combines novelty search with evolution. Instead of evolving a single optimal genome, we want to evolve an optimal genome for every point in some grid of features.
In our case, we can use the two metrics from before – node count and bond ratio. Every spot in the matrix corresponds to some range of the metrics (e.g. 50-60 nodes and 30%-40% bonds), and we store the optimal genome that fits the criteria.
MAP-Elites works by:
- Maintain a 32 x 32 matrix of genomes.
- Pick a random genome, and mutate it.
- Measure its metrics, and find which spot in the grid the genome belongs to.
- If the new genome performs better than the old genome in the spot, replace it.
As always, let's get our hands dirty and check out what the optimal genomes end up looking like.
In the MAP-Elites genomes, we get a whole bunch of different creatures, and some cool patterns emerge. The fastest creatures turn out to be the smaller ones, likely due to their lower mass. Larger creatures tend to develop a kind of "tail" -- my guess is that this acts as a stabilizer so the creature doesn't drift off direction.
The last row is especially interesting, since the larger creatures develop a completely different movement strategy. Instead of moving its entire body, the creature unrolls itself to stretch forwards.
Quality-diversity algorithms are an easy way to explore the possibilty space of an open-ended world. The techniques, such as novelty search and MAP-Elites, are simple but yield strong results when given proper metrics of diversity. In this Oceanworld, the limiting factor is the expressivity of our genome -- there are only so many ways to build swimming creatures out of a single type of bond.
Further work can involve:
- How can we increase the expressivity of our genome? Can we make the genome more information-dense in relation to the creature it defines? What kinds of genomes allow innovations to transfer between creatures?
- What are good definitions of diversity? Can we define diversity in terms of a creature's behavior, rather than its structure?
In the MAP-Elites method, it turns to be helpful to first run novelty search to get a population of diverse genomes. We start the MAP-Elites matrix with this diverse population, rather than a population of random genomes.
MAP-Elites from random starting population.
MAP-Elites from novelty-search population.
For more on quality-diversity, a good starting point are the ICML tutorial slides below. The rest are some highlight papers with useful ideas.
"Recent Advances in Population-Based Search", ICML 2019 Tutorial, by Jeff Clune, Joel Lehman, and Ken Stanley. This is a great presentation that goes over recent works related to quality diversity. It introduces novelty search as a way to get around environments where greedily optimzing traps you in a local minimum, and then goes to to motivate quality-diversity algorithms where we simultaneously train populations of high-performing agents.
"Abandoning Objectives: Evolution through the Search for Novelty Alone", by Joel Lehman and Ken Stanley. This is the paper on Novelty Search. It argues that basing evolutionary search on fitness is limiting, since only creatures with high fitness will be considered. In environments such as hard mazes, the fitness landscape is non-convex and greedy optimization will fail. Instead, strong diversity metrics are often a better landscape to search through.
"Illuminating search spaces by mapping elites", by Jean-Baptiste Mouret and Jeff Clune. This is the MAP-Elites paper. They introduce the idea of "illumination algorithms", which aim to find the highest-performing solution for every point in a given feature space. The core MAP-Elites idea is pretty straightforward -- store the best solution at every point, and keep trying new solutions.
"POET: Paired Open-Ended Trailblazer", by Rui Wang et al. POET is a quality-diversity algorithm with a step towards open-endedness. The idea is similar to MAP-Elites, but instead of niches defined as properties of the genome, each niche is a task. New tasks are introduced randomly, and stay in the pool if agents can solve them. You end up with a growing population of tasks and population of agents that can solve those tasks.
What are good diversity metrics? Hand-designed metrics such as creature size only work to some degree. In my mind, this is an important question as "novelty" is hard to define concretely.
"Diversity is All You Need", by Benjamin Eysenbach et al. This paper describes a system where we want to find a set of diverse policies. They define diversity as:
- A discriminator should be able to predict which policy is active based on state history.
- A policy should cover as many states as it can.
"Stochastic Neural Networks for Hierarchical Reinforcement Learning", by Carlos Florensa, Yan Duan, and Pieter Abeel. Motivated by information theory, this work aims to find diverse policies by maximizing entropy. This ends up being a similar method as above -- policies should visit different states to be maximally diverse.
"Autonomous skill discovery with Quality-Diversity and Unsupervised Descriptors", by Antoine Cully. This work takes hints from dimensionality reducing methods such as PCA (Principle-component analysis). Basically, given a set of trajectories, we can map every trajectory onto a 2D space. This mapping becomes our diversity metric for running a quality-diversity method such as MAP-Elites.
Aside from how to discover diverse populations, the world and environment this all takes place in is crucial as well. These are some nice references for work in ALIFE and evolved creatures.
"Evolution Strategies", OpenAI. A good overview of how evolutionary systems can potentially outscale traditional reinforcement learning. The basic idea is that ES is inherently parallalizable, so systems with lots of CPU cores can scale well. ES also treats an entire episode with a single reward, so it doesn't suffer from things such as sparsity or time dependency.
"Evolving Virtual Creatures", by Karl Sims. This is a classic work from 1994, where virtual swimming robots are evolved to move forwards. The methods used as similar to what we did in this blog post, but the genome is defined as a hierarchy of parts, and there are basic neural networks for control.
"Evolving soft locomotion", by Francesco Corucci et al. A more recent paper that uses evolutionary methods to design soft swimming and walking robots. The world here uses creatures of deformable voxels, akin to legos, that apply a fluid resistance force when expanding (similar to what we did but in 3D). Their genome is a CPPN, a neural network C(x,y) that outputs the creature's properties at each coordinate.
Follow @kvfrans